 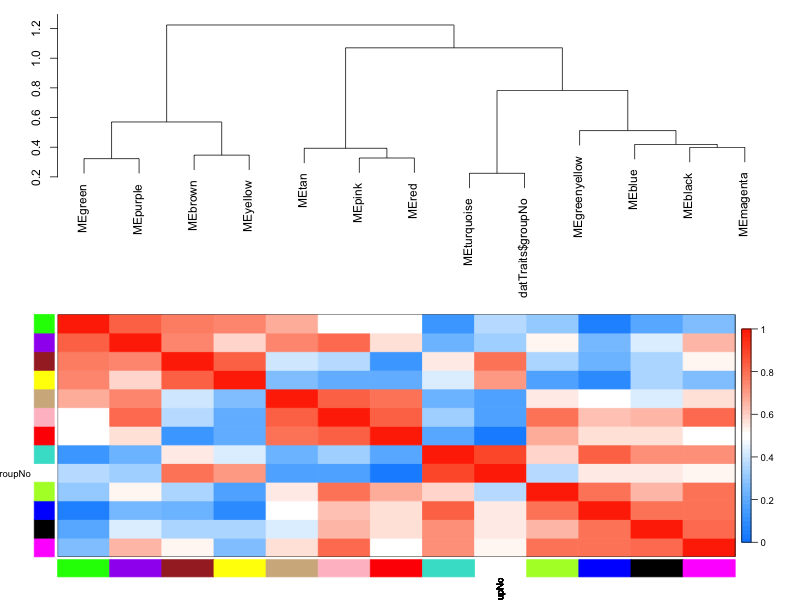
I've also run the analysis from scratch to avoid loading the TOM, and I get the same error. However, following the advice in that bioconductor thread leads to another error: TOM.mat = as.matrix(TOM)Įrror in TOM.mat : (subscript) logical subscript too long Previously it was suggested that this error might be related to loading the TOM from file, in which case it is not saved as a proper similarity matrix ( ). This function exports a network in edge and node list files in a format suitable for importing to Cytoscape. The co-expression network was constructed by selecting the genes whose variance was greater than all the quartiles of variance. Note that a minimum threshold of 0.05 should always be applied in order to reduce the size of the Edge- and Node-files. 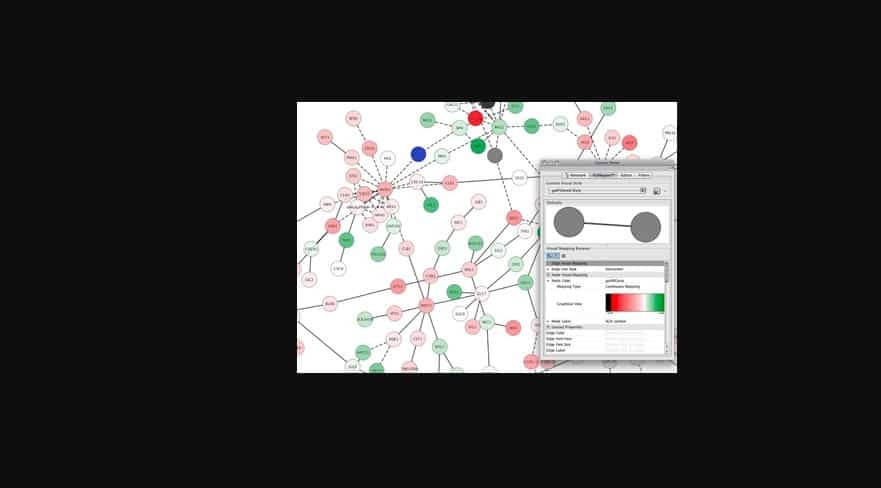
# Select the corresponding Topological OverlapĮrror in TOM : incorrect number of dimensions WGCNA is a widely used systems biology method that is usually used to establish a scale-free network based on gene expression data profiles 18. load (file 'Data/WGCNAcourseNet1adjacencymatrix.RData') adjacency-treshold, genes that are connected with with edge-weights lower than this threshold will be excluded from the Edge- and Node-files. The steps in R are as follows: # choose module I should note here that I used the blockwiseModules function to create the TOM because of the size of my RNAseq data set. I only have gene expression data, fpkm, and NO trait data. Tutorial 6 on the WGCNA page deals with this. I am doing the gene co-expression network using WGCNA, and want to export the network to Cytoscape so that I can see modules in Cytoscape. This step requires the topological overlap matrix (TOM) to be subset. The key parameters were set as follows: soft threshold power, 12 min module size, 20 merge cut height, 0.25. 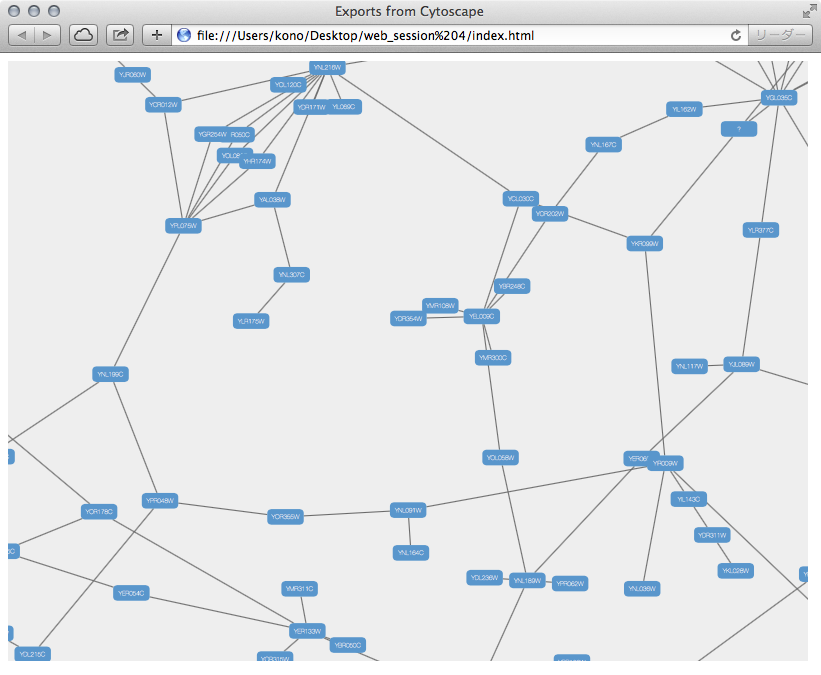
I am trying to export my results from WGCNA for network visualisation with visANT or Cytoscape. WGCNA The 1033 candidate genes involved in the SS biosynthesis pathway were the input of R package WGCN, and a weighted co-expression network was built to reveal potential saponin biosynthesis modules of B. 
0 Comments
Leave a Reply. |
 RSS Feed
RSS Feed
